Rapid tissue oxygenation mapping from snapshot structured-light images with adversarial deep learning
Chen MT, Durr NJ
Journal of Biomedical Optics 25:11 2020
[Journal Link] [Editor’s Spotlight] [Cover Article] [Preprint]
Abstract
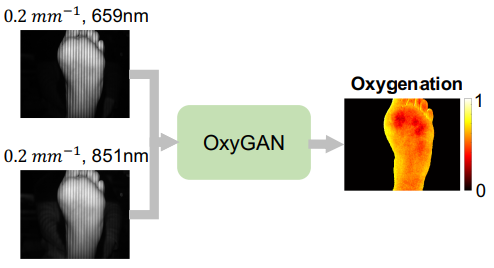
Spatial frequency domain imaging (SFDI) is a powerful technique for mapping tissue oxygen saturation over a wide field of view. However, current SFDI methods either require a sequence of several images with different illumination patterns or, in the case of single snapshot optical properties (SSOP), introduce artifacts and sacrifice accuracy. To avoid this tradeoff, we introduce OxyGAN: a data-driven, content-aware method to estimate tissue oxygenation directly from single structured light images using end-to-end generative adversarial networks. Conventional SFDI is used to obtain ground truth tissue oxygenation maps for ex vivo human esophagi, in vivo hands and feet, and an in vivo pig colon sample under 659 nm and 851 nm sinusoidal illumination. We benchmark OxyGAN by comparing to SSOP and to a two-step hybrid technique that uses a previously-developed deep learning model to predict optical properties followed by a physical model to calculate tissue oxygenation. When tested on human feet, a cross-validated OxyGAN maps tissue oxygenation with an accuracy of 96.5%. When applied to sample types not included in the training set, such as human hands and pig colon, OxyGAN achieves a 93.0% accuracy, demonstrating robustness to various tissue types. On average, OxyGAN outperforms SSOP and a hybrid model in estimating tissue oxygenation by 24.9% and 24.7%, respectively. Lastly, we optimize OxyGAN inference so that oxygenation maps are computed ~10 times faster than previous work, enabling video-rate, 25Hz imaging. Due to its rapid acquisition and processing speed, OxyGAN has the potential to enable real-time, high-fidelity tissue oxygenation mapping that may be useful for many clinical applications.